Ethan's BGGN 213 Portfolio
Class work for bioinformatics
HW06
Ethan Ashley (PID: A15939817) 2025-10-17
Generalize the following protein analysis code to work for any protein sequence
# Can you improve this analysis code?
library(bio3d)
s1 <- read.pdb("4AKE") # kinase with drug
## Note: Accessing on-line PDB file
s2 <- read.pdb("1AKE") # kinase no drug
## Note: Accessing on-line PDB file
## PDB has ALT records, taking A only, rm.alt=TRUE
s3 <- read.pdb("1E4Y") # kinase with drug
## Note: Accessing on-line PDB file
s1.chainA <- trim.pdb(s1, chain="A", elety="CA")
s2.chainA <- trim.pdb(s2, chain="A", elety="CA")
s3.chainA <- trim.pdb(s1, chain="A", elety="CA")
s1.b <- s1.chainA$atom$b
s2.b <- s2.chainA$atom$b
s3.b <- s3.chainA$atom$b
plotb3(s1.b, sse=s1.chainA, typ="l", ylab="Bfactor")
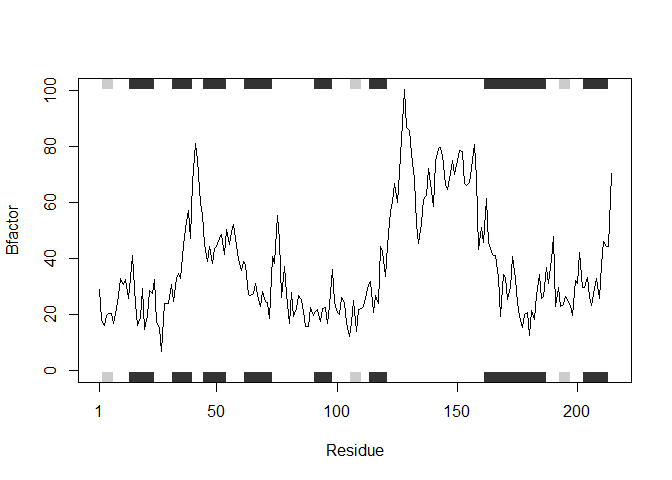
plotb3(s2.b, sse=s2.chainA, typ="l", ylab="Bfactor")
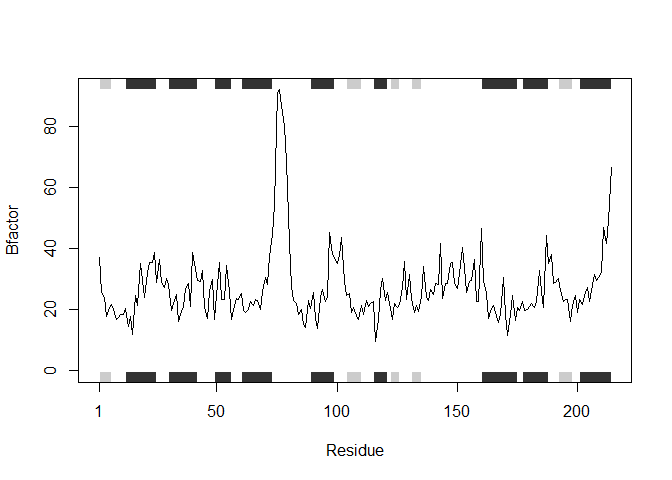
plotb3(s3.b, sse=s3.chainA, typ="l", ylab="Bfactor")
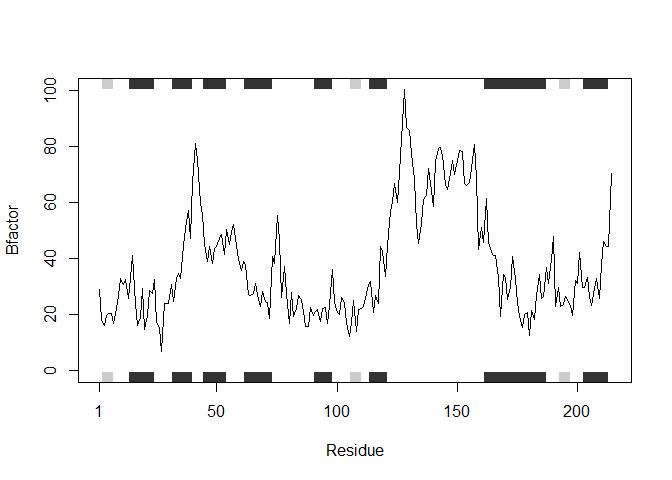
Q6. How would you generalize the original code above to work with any set of input protein structures?
Converting the general code above into a more generalized version
inputProteins = c("4AKE", "1AKE", "1E4Y")
#generating a for loop that takes a list of PDB protein ID's and analyzes each one individually as shown above
for (i in inputProteins) {
s <- read.pdb(i)
s.chainA <- trim.pdb(s, chain="A", elety="CA")
s.b <- s.chainA$atom$b
plotb3(s.b, sse=s.chainA, typ="l", ylab="Bfactor")
}
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/4AKE.pdb exists. Skipping download
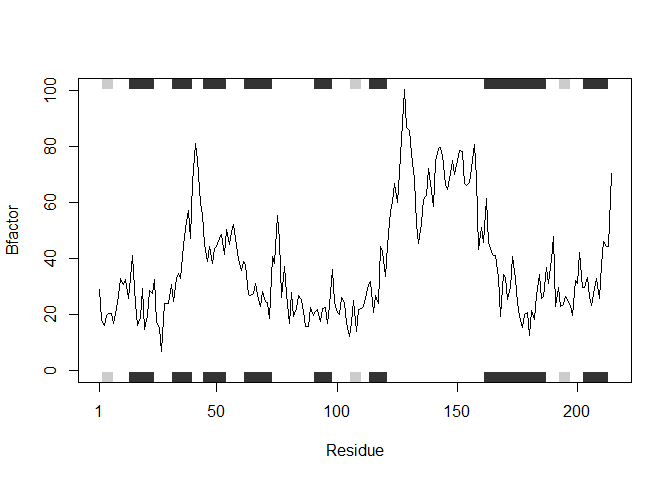
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1AKE.pdb exists. Skipping download
## PDB has ALT records, taking A only, rm.alt=TRUE
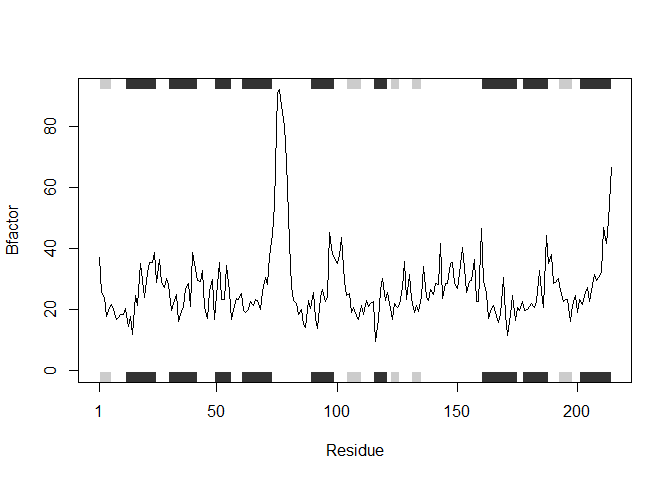
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1E4Y.pdb exists. Skipping download
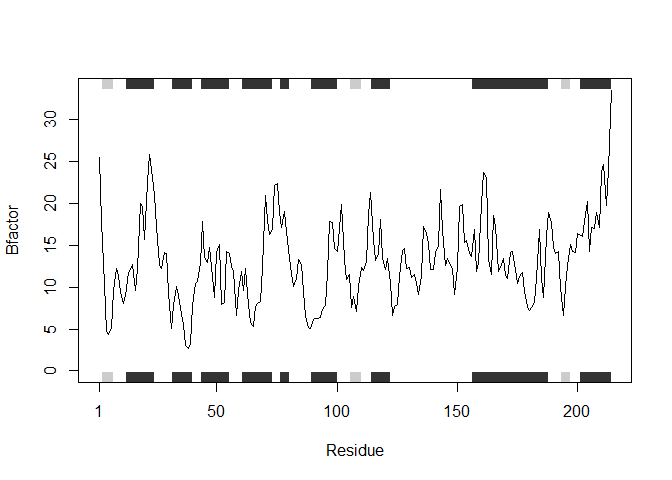
Converting the generalized code into a function that can be applied to any list of sequences
#This function accepts a list of PDB protein ID's and analyzes each one individually as shown above. main is added as an input to plotb3 to clarify which protein is plotted.
analyzeProteins <- function(seqs) {
for (i in inputProteins) {
s <- read.pdb(i)
s.chainA <- trim.pdb(s, chain="A", elety="CA")
s.b <- s.chainA$atom$b
plotb3(s.b, sse=s.chainA, typ="l", ylab="Bfactor", main=i)
}
}
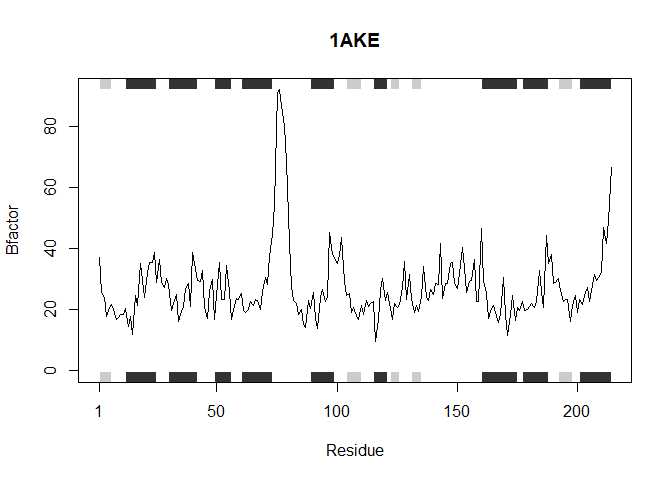
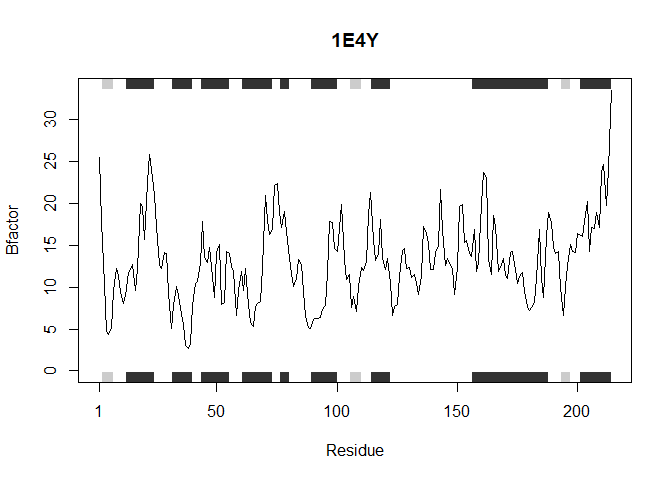
This function accepts a list of PDB protein ID’s and analyzes each one individually as shown above generating a plot of the residue position vs. the B factor
Testing new function on the provided list of sequences
analyzeProteins(inputProteins)
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/4AKE.pdb exists. Skipping download
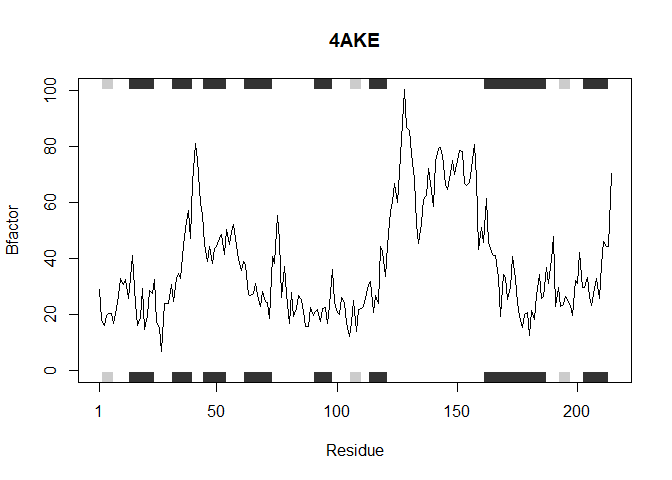
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1AKE.pdb exists. Skipping download
## PDB has ALT records, taking A only, rm.alt=TRUE
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1E4Y.pdb exists. Skipping download
Testing with sapply()
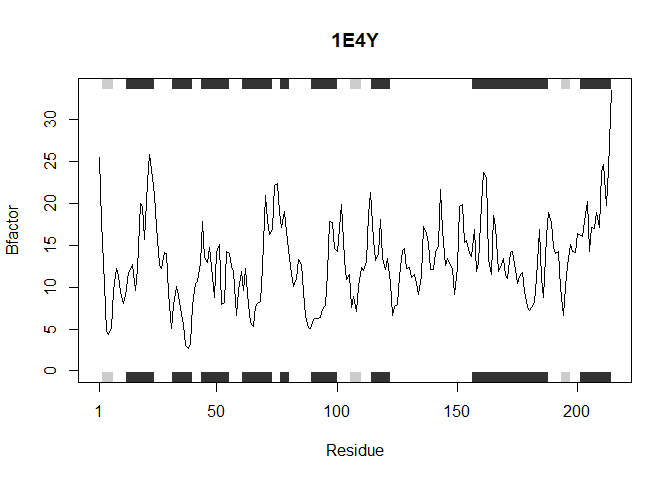
Making the function even more generalized so that it can be utilized with sapply(). It accepts a single PDB ID at a time but can be vectorized with sapply() to achieve the same output as previously
analyzeProteins <- function(seqs) {
s <- read.pdb(i)
s.chainA <- trim.pdb(s, chain="A", elety="CA")
s.b <- s.chainA$atom$b
plotb3(s.b, sse=s.chainA, typ="l", ylab="Bfactor", , main=i)
}
sapply(inputProteins, analyzeProteins)
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1E4Y.pdb exists. Skipping download
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1E4Y.pdb exists. Skipping download
## Note: Accessing on-line PDB file
## Warning in get.pdb(file, path = tempdir(), verbose = FALSE):
## C:\Users\eashl\AppData\Local\Temp\RtmpolJCYC/1E4Y.pdb exists. Skipping download
## $`4AKE`
## NULL
##
## $`1AKE`
## NULL
##
## $`1E4Y`
## NULL